SO-Pharm: OBO Foundry requirments
v2.0 alpha (2007/08/21) |
|
|
|
|
This page reports how SO-Pharm answers to
the 10 OBO Foundry Principles
1. The ontology must be open and available to be used by all without any
constraint other than (a) its origin must be acknowledged and (b) it is not to
be altered and subsequently redistributed under the original name or with the
same identifiers.
2. The ontology is in, or can be expressed in, a common shared syntax. This
may be either the OBO syntax, extensions of this syntax, or OWL.
>>>> SO-Pharm is expressed in OWL-DL.
>>>> See the WonderWeb OWL Ontology Validator at
>>>> http://www.mygrid.org.uk/OWL/Validator
3. The ontologies possesses a unique identifier space within the OBO Foundry.
>>>> The unique prefix of SO-Pharm is SOPHARM which is related to the namespace
>>>> "http://www.loria.fr/~coulet/ontology/sopharm/version2.0/sopharm.owl#"
>>>> A persistent URL is http://purl.oclc.org/NET/loria/sopharm/owl/sopharm
>>>> is available.
>>>> Each class and property is identified by a unique identifier of the form
>>>> SOPHARM:SOPHARM_00100, the name of the class (or property) could be found
>>>> in the section "label" of meta-data.
4. The ontology provider has procedures for identifying distinct successive
versions.
>>>> The unique namespace of the ontology depends on its version (see 3.).
>>>> The version of the ontology is also displayed in OWL meta-data.
>>>>
>>>> >Author: Adrien Coulet
>>>> Version: SO-Pharm 2.0
>>>> Release: 07/18/2007
>>>> Contact: adrien dot coulet at loria dot fr
>>>> University of Nancy
5. The ontology has a clearly specified and clearly delineated content.
The ontology must be orthogonal to other ontologies already lodged within
OBO.
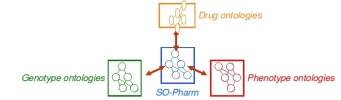
>>>> SO-Pharm is a Suggested Ontology for Pharmacogenomics.
>>>> It is a formal representation of domain knowledge in pharmacogenomics.
>>>> SO-Pharm articulates ontologies from sub-domains of pharmacogenomics
>>>> (i.e. genotype, phenotype, drug, clinical trials).
>>>> It enables the representation of pharmacogenomic relationships between drugs,
>>>> genomic variations and phenotype traits.
>>>> It enables the management of knowledge for case-study, investigation and
>>>> test over new hypothesis in pharmacogenomics.
>>>> SO-Pharm was specially developed because of the lack of available ontology
>>>> that overlaps sub-domains of pharmacogenomics.
6. The ontologies include textual definitions for all terms.
>>>> Every class and property of SO-Pharm has a textual definition.
>>>> The used language is english.
7. The ontology uses relations which are unambiguously defined following the
pattern of definitions laid down in the OBO Relation Ontology.
>>>> Object properties of SO-Pharm are mapped, when possible, to properties from
>>>> the Relationship Ontology (OBO_REL).
8. The ontology is well documented.
>>>> Information about modifications, versioning, future works in SO-Pharm is
>>>> available in OWL meta-data.
>>>> Meta-data, organisation of the ontology and much more are documented in
>>>> OWLDoc available at
>>>> http://www.loria.fr/~coulet/ontology/sopharm/version2.0/owldoc/index.html
>>>> Additional informations about SO-Pharm construction and validation are
>>>> detailed in
>>>> Coulet, A.et al.: Suggested Ontology for Pharmacogenomics (SO-Pharm):
>>>> Modular Construction and Preliminary Testing. Springer, LNCS 4277: 648-57.
9. The ontology has a plurality of independent users.
>>>> Not discussed here.
>>>> The ontology is used in several collaborations. Contact Adrien Coulet
>>>> (adrien dot coulet at loria dot fr) for more informations.
10. The ontology will be developed collaboratively with other OBO Foundry
members.
>>>> We are working on identifying potential overlaps of SO-Pharm with other
>>>> ontologies to collaborate with related people and groups (e.g. PharmGKB group).
>>>> In addition, we are in constant dicussion with Pierre Grenion from the Institute
>>>> for Formal Ontology and Medical Information Science at Saarland University
Author
Adrien Coulet
Contact
adrien dot coulet at loria dot fr
http://www.loria.fr/~coulet
University of Nancy

