-
-
In Multiple Instance Learning (MIL) problem for sequence data, the learning data consist of a set of bags where each bag contains a set of instances/sequences. In many real world applications such as bioinformatics, web mining, and text mining, comparing a random couple of sequences makes no sense. In fact, each instance of each bag may have structural and/or temporal relation with other instances in other bags. Thus, the classification task should take into account the relation between semantically related instances across bags. In this paper, we present two novel MIL approaches for sequence data classification: (1) ABClass and (2) ABSim.
We applied both approaches to the problem of bacterial Ionizing Radiation Resistance (IRR) prediction. We evaluated and discussed the proposed approaches on well known Ionizing Radiation Resistance Bacteria (IRRB) and Ionizing Radiation Sensitive Bacteria (IRSB) represented by primary structure of basal DNA repair proteins. The experimental results show that both ABClass and ABSim approaches are efficient.
- Naive approach: The naive approach for MIL in sequence data consists of a two step approach. The first step is a preprocessing step that transforms the set of sequences to an attribute-value matrix where each row corresponds to a sequence and each column corresponds to an attribute. The second step consists in applying an existing MIL classifier. It is worthwhile to mention that only a subset of the used attributes is representative for each processed sequence. Therefore, we may have a big sparse matrix when trying to present the whole sequence data using an attribute value format.
- ABClass approach: In order to avoid the use of one large vector of features to describe sequence data, we present ABClass, a novel approach that takes into account the across bag relations. Each set of related instances will be presented by its own motifs vector. This reduces the number of attributes that are not representative for the processed sequence. Instead of using a classifier that uses a large vector to describe all the sequences data, every vector of motifs will be used to produce a prediction result. These results will be then aggregated to have a final result.
- ABSim approach: According to the specificity of the processed data, a similarity measure can be defined and used to discriminate instances. ABSim focuses on discriminating bags by measuring the similarity between each instance sequence in the query bag and corresponding related sequences in the different bags of the learning database.
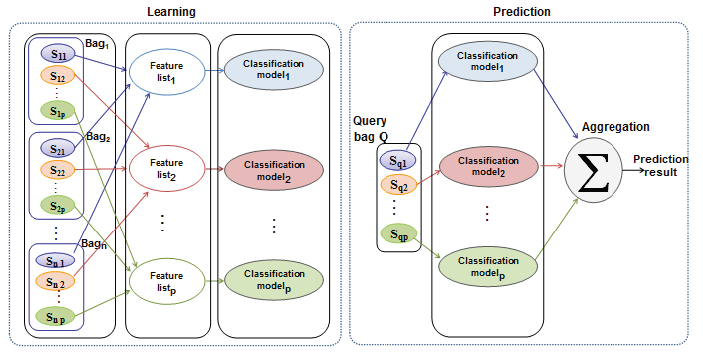
Fig 1. System overview of the ABClass approach
- General description
- We evaluated and discussed the proposed approaches on well known Ionizing Radiation Resistance Bacteria (IRRB) and Ionizing Radiation Sensitive Bacteria (IRSB) represented by primary structure of basal DNA repair proteins. We constructed a database containing 14 IRRB and 14 IRSB. Each bacterium contains 25 to 31 proteins implicated in basal DNA repair in IRRB.
- Data source
- Proteins of the bacterium Deinococcus radiodurans were downloaded from the UniProt web site. http://www.uniprot.org/uniprot/
- PerfectBlast tool was used to identify orthologous proteins of the others bacteria. (Tool downloadable here)
- Proteomes of other bacteria were downloaded from the NCBI FTP web site. http://www.ncbi.nlm.nih.gov/Ftp/
-
Computations were carried out on a i7 CPU 2.49 GHz PC with 6 GB memory, operating on Linux Ubuntu. In the classification process, we used the Leave-One-Out (LOO) technique.
Both ABClass and ABSim approaches provide good overall accuracy results since the least accuracy percentage is 89.2%. This clearly shows that our proposed approaches are efficient. Using ABSim approach with the SMS aggregation method provides a better accuracy result compared to the WAMS aggregation method. The best result was reached using ABClass approach, J48 classifier and the motif extraction settings 3 and 4. Using these two settings, a large number of non discriminative motifs are extracted.
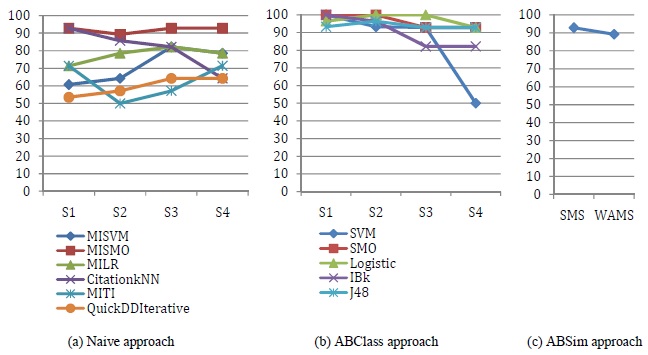
Fig 2. Accuracy percentage using the naive approach, ABClass approach and ABSim approach.
- ABClass implementation
- ABClass implementation runs on a Windows or a Linux platform (tested on Ubuntu distribution) that contains a java JRE. - Version: 2.0 ( May - 2019) ABClass for Windows 64 bit is downloadable here. - Version: 1.0 ABClass for Windows 64 bit is downloadable here. ABClass for Linux 64 bit is downloadable here. You can download the dataset used in our experiments here.
- ABSim implementation
- ABSim implementation runs on a Windows or a Linux platform (tested on Ubuntu distribution) that contains a java JRE. ABSim for Windows 64 bit/Linux 64 bit is downloadable here. You can download the dataset used in our experiments here.
-