Targeting a protein – DNA complex to inhibit the transfer of antimicrobial resistance
Regional project (Mirabelle+ and FEDER) N. Leblond‐Bourget. Collab DynAMic lab (UL/INRA) & LPCT lab (UL/CNRS)
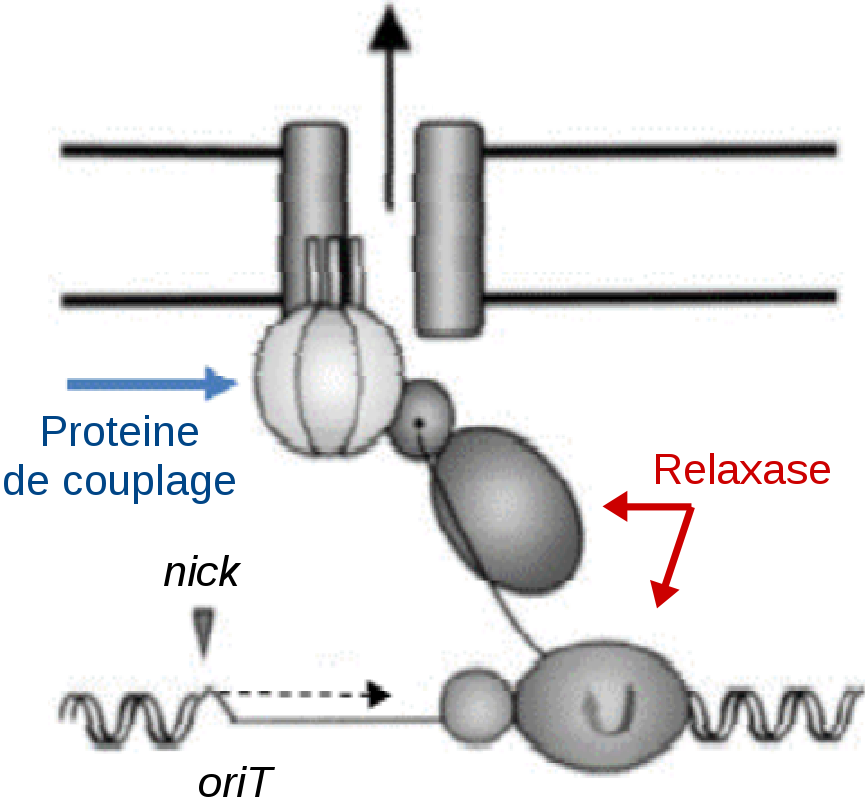
The transfer of resistance to antibacterials often results from the transfer of mobile genetic elements _ such as ICEs _ by conjugation. ICEs encode in particular a conjugation trans-membrane pore allowing their own transfer to a recipient cell. The CITRAM project aims to develop conjugation inhibitors by targeting relaxase. This key protein of the conjugation (i) recognizes and cleaves the DNA to be transferred, (ii) binds with this DNA to form a “relaxosome” and (iii) interacts with a coupling protein that allows the transfer of the DNA via the conjugation pore. We apply to a model ICE combined approaches of molecular bioengineering, biochemistry, structural bioinformatics, virtual screening and molecular simulation to design, and model potential inhibitors of relaxase at these three levels.
This project includes methodological developments: 1/ the expansion of our protein-ssRNA binding method to dock DNA with both single/double-stranded regions, and 2/the use of various experimental data from biochemistry and biophysics to guide the docking, i.e. data-driven / integrative modeling.



