Software to the development of which I contributed
ATTRACT: macromolecular docking
ATTRACT is a macromolecular docking program : it predicts the three-dimensional structure of complexes made of proteins, peptides and/or nucleic acids (technical details below). ATTRACT program is free and open-source, and applicable to protein – protein/peptides/nucleic acids complexes. The usage of its many options is eased for people outside bio-informatics by an ATTRACT web-interface www.attract.ph.tum.de. The interface provides guidance and feedback on the various options and parameters, and performs extensive validation of the user input [De Vries 2015].
Docking with ATTRACT starts with placing the ligand at or randomly initial positions around the receptor. The 3 positional and 3 rotational degrees of freedom are then explored by a series of minimisations of the intermolecular energy, computed as the sum of atom-atom interaction energy. Computation can be made extremely fast by two features. First, a coarse grained representation groups atoms of each residue into 3 or 4 beads, and bead-bead energies are defined by a knowledge-based force-field [Zacharias 2005, Setny 2011]. Second, a pre-calculated grid stores the interaction energy for each possible bead with the receptor, speeding up the calculation by a factor 30-100 [De Vries 2017]. The final poses are re-scored, using either knowledge-based or step potentials [Sasse 2017]. The coarse-grained representation accounts for flexibility of small amplitudes (e.g. side-chains conformations). Options are available to account for larger conformational changes between bound and unbound molecules, for both receptor and ligand: (i) normal modes can be pre-computed and added as additional degrees of freedom, (ii) ensembles of pre-computed conformations can be docked in parallel, (iii) domains with variable respective positions due to flexible linkers can be docked as separate bodies with distance restraints accounting for the possible linker conformations, (iv) final refinement of poses can be performed with partial or full flexibility. Experimental data on the complex to be modelled can also be added as distance restraints. MONETA provides an atomic-scale structural description of allosteric communications in proteins or nucleic acids. It analyses molecular dynamics simulations to extract dynamics correlations between atoms, and extracts (i) spacial clusters of highly coupled residues, and (ii) chains of residues as communication pathways connecting those clusters [Laine 2011]. The communication network can be visualised by interactive representations on the 3D structure of the macromolecule (PyMOL module) or on a 2D map of connected residues (GEPHI module) [Allain 2014]. Those visualisation tools can ease for instance the detection of communication hubs, or the understanding/prediction of the impact of a mutation by comparison of wild-type vs mutant communication profiles. We applied MONETA to analyse and compare the communication pathways (i) in the native vs mutant forms receptors tyrosine kinases KIT [Chauvot de Beauchene 2016] and CSF-1R [D.S.F.C. Gomes 2016;] , and (ii) in the phosphorylated vs non-phosphorylated proteins STAT5 [Langenfeld 2015]. These studies shed light on (a) the constitutive activation induced by equivalent mutations in KIT and CSF-1R, and (b) allosteric regulation in the activated and non-activated STAT5 proteins.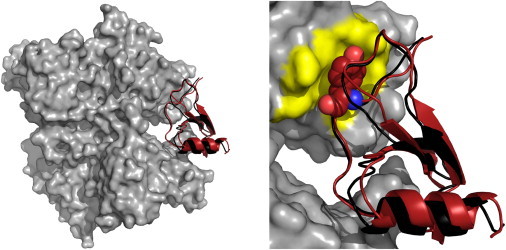
MONETA : Allosteric communication



